A modern workbench for
molecular biology.
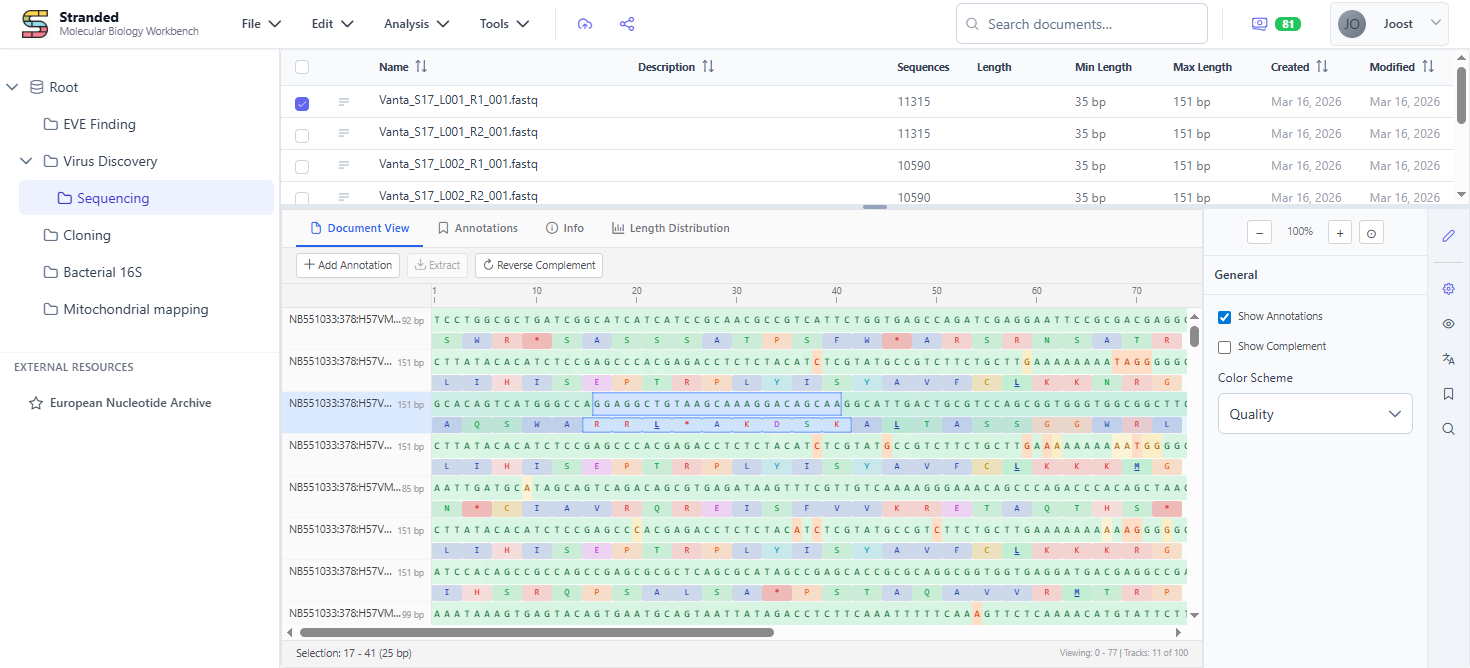
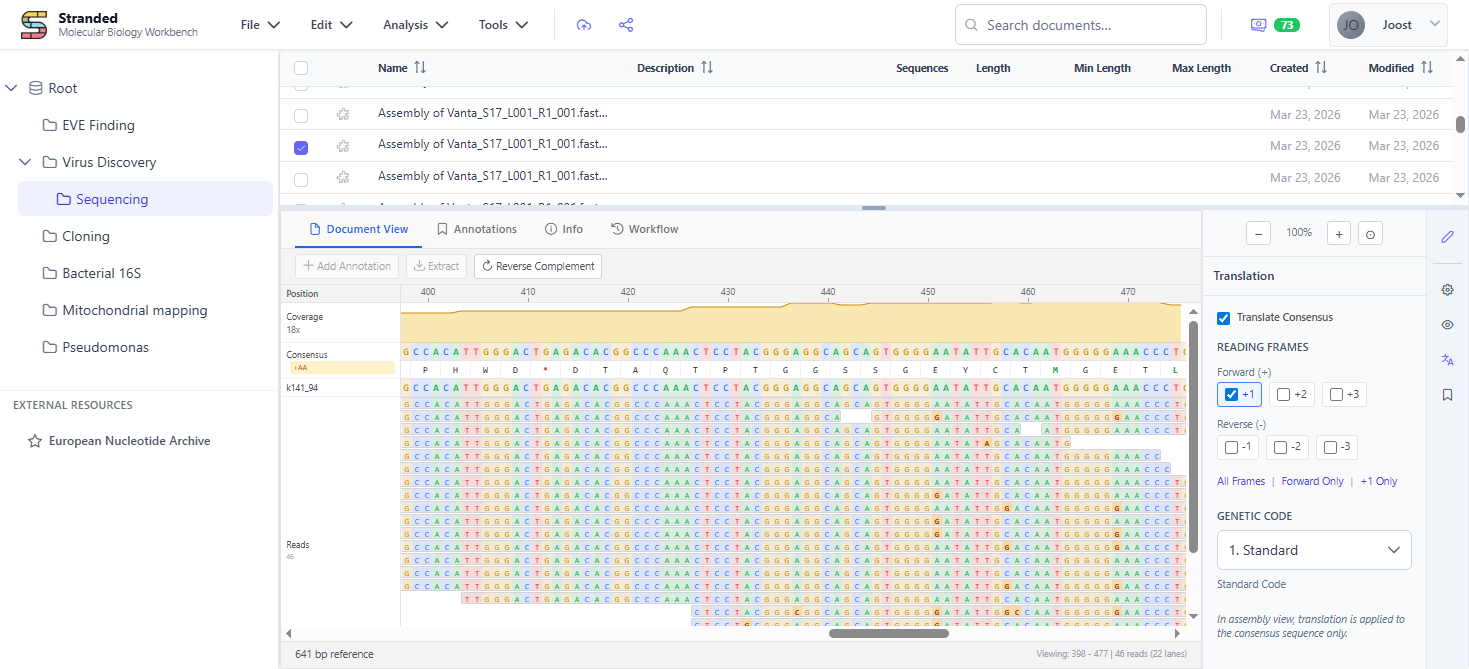
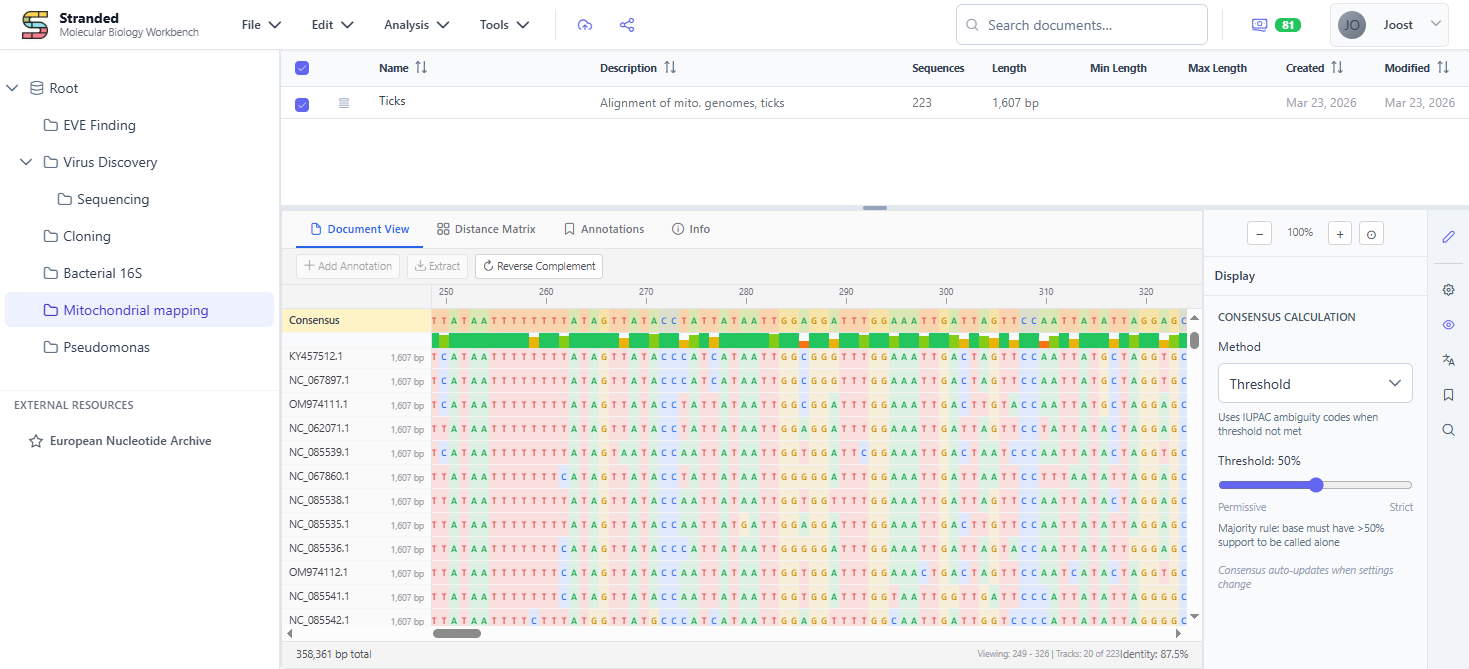
Stranded is a browser-based platform for working with sequences, chromatograms, alignments, assemblies and phylogenies. One central spot for all your molecular work - import, annotate, analyse, and share without switching tools.
Made & hosted in the EU, and GDPR compliant!
Academic-first onboarding · Q2 2026